Beleggia Lab
Functional cancer genomics
Small Cell Lung Cancer (SCLC) is one of the most aggressive types of cancer, with a 5-year survival rate of 3% for patients diagnosed in advanced stage and 16% for patients diagnosed at an early stage. Despite intensive efforts, only minor advances in the pharmacological therapy of SCLC have been produced in the last decades: no targeted therapy has entered the clinic and immunotherapy only provides a marginal improvement.
Our research focus:
The focus of our research is the integration of different large-scale OMICS data, both generated within the lab and available in the literature, in order to understand the differences between cancer cells and healthy cells.
Our goals:
The aim of our research is to characterize the defining molecular and phenotypic characteristics of SCLC and investigate how they can be exploited to improve the therapy of SCLC.
Our approach:
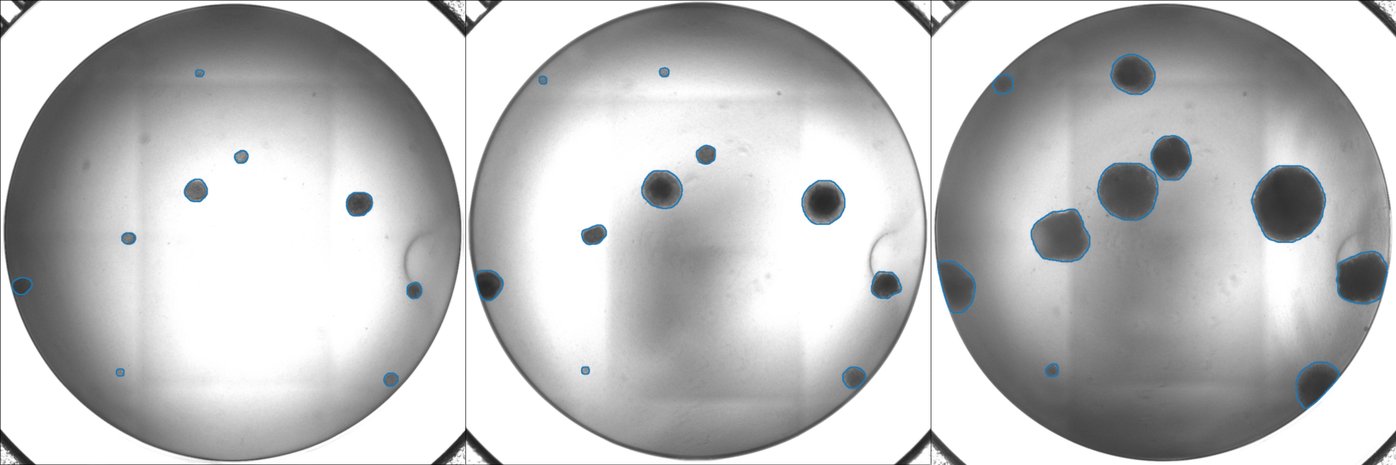
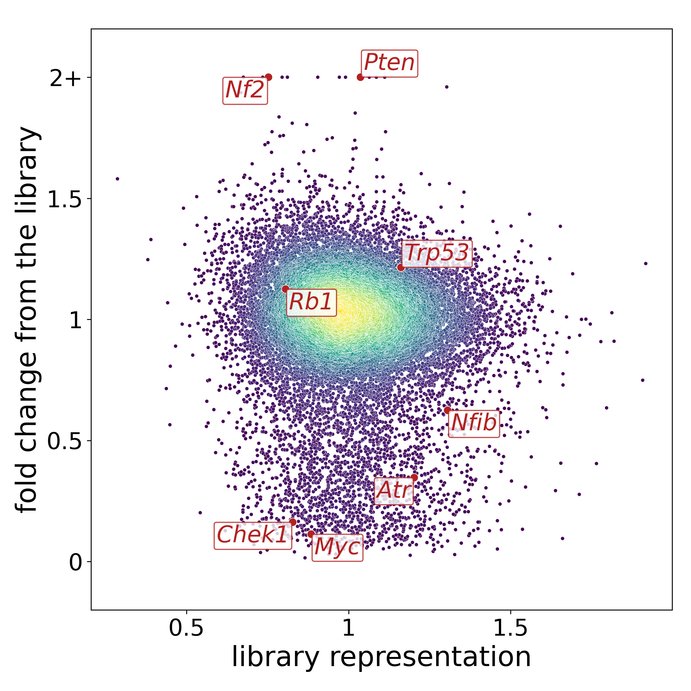
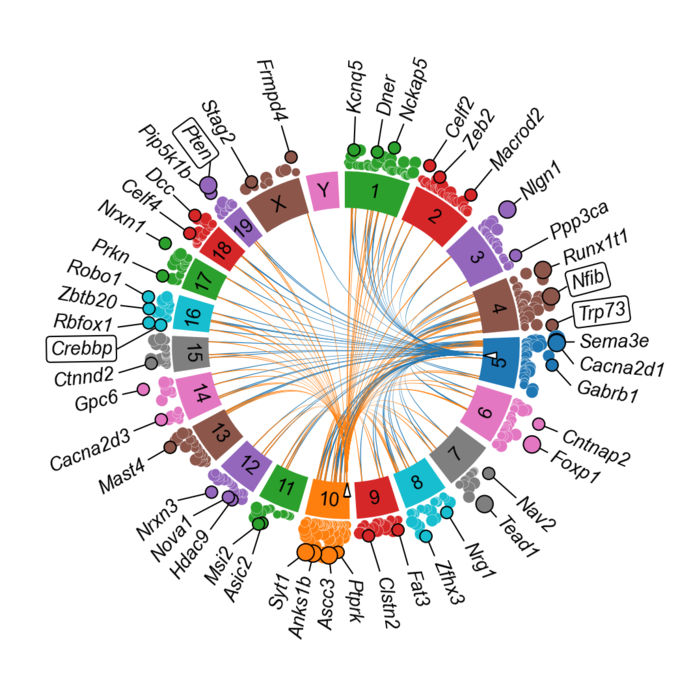
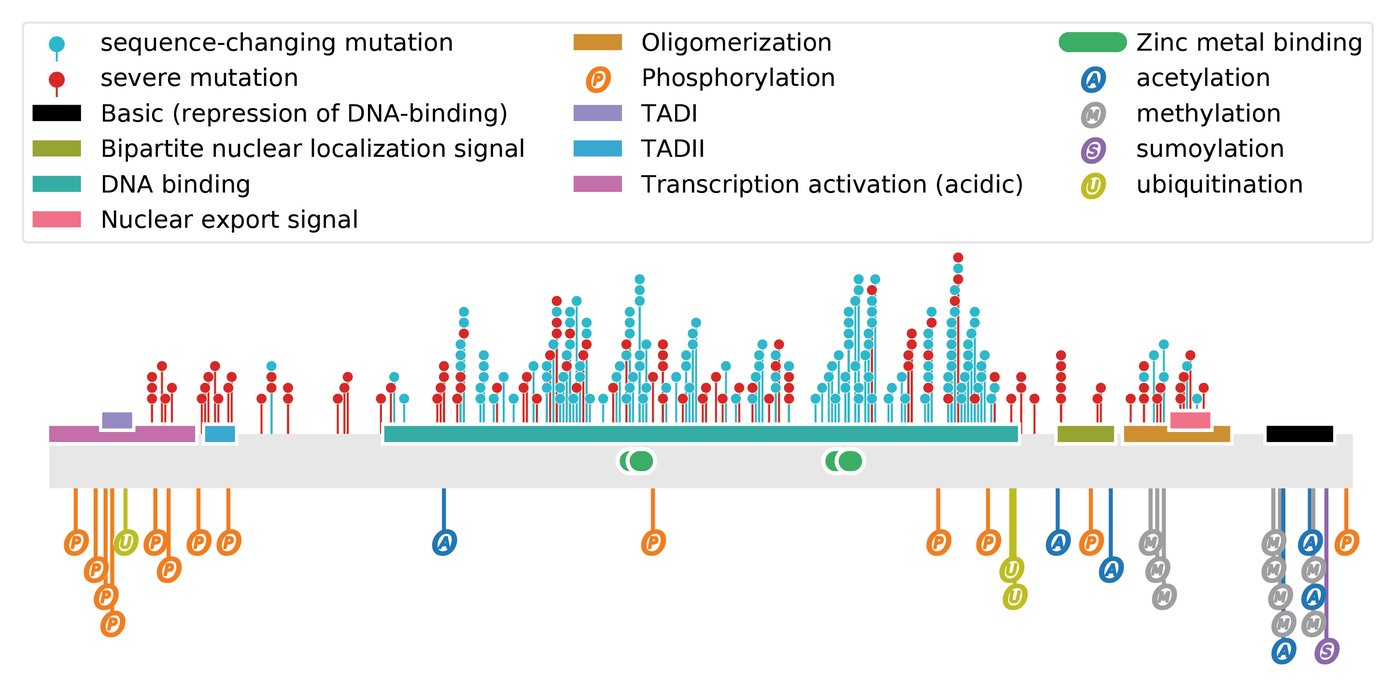
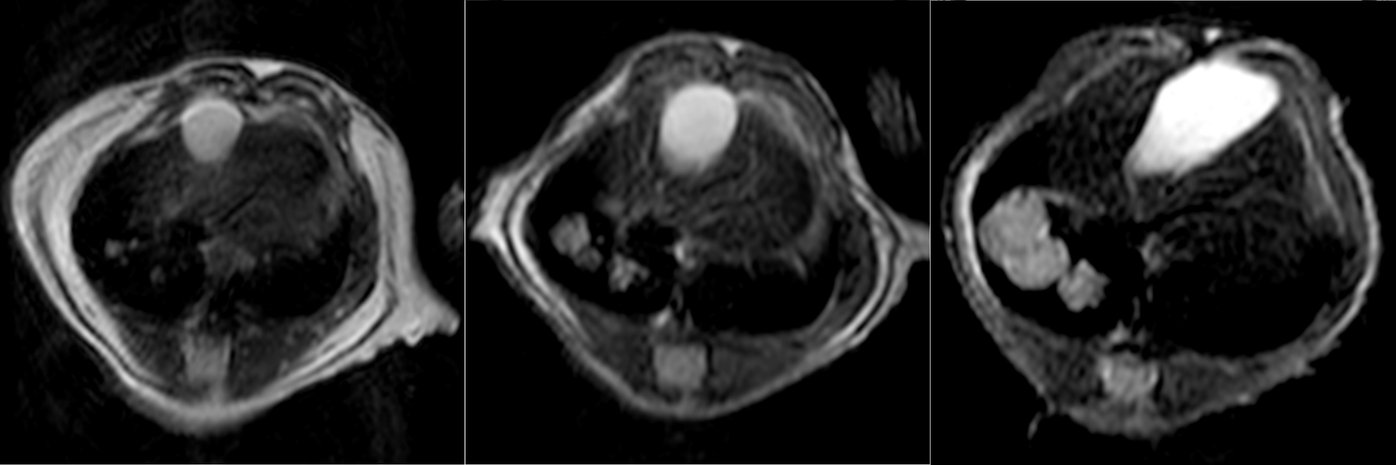
In an effort to obtain a solid and unbiased overview of the phenotype of SCLC, that can support the translation into actionable therapeutic targets, we use large-scale genetic tools, such as whole-genome CRISPR-Cas9 screens (Fig. 1) and in vivo screens (Fig. 2) as well as data from patient samples and comprehensive online databases (Fig. 3). We then use autochthonous mice (Fig. 4) and organoid cultures (Fig.5) to test novel therapeutic strategies in the first line and second line setting.
Principal Investigator
Team

Maike Boecker
Student

Manoela Caiaffa
Student

Ali Chitsaz
MD Student
Most important publications
- Schmitt A, Sakthivelu V, Ndoci K, Wani GA, Touet M, Pintelon I, Kisis I, Ibruli O, Weber J, Maresch R, Bebber CM, Goergens J, Jevtic M, Odenthal F, Placzek A, Hennrich AA, Conzelmann KK, Boecker M, Heimsoeth A, Gülcüler GS, Jachimowicz RD, George J, Brägelmann J, von Karstedt S, Peifer M, Persigehl T, Grüll H, Sos ML, Brüning J, Reifenberger G, Fischer M, Adriaensen D, Büttner R, Brouns I, Rad R, Thomas RK, Bergami M, Motori E, Reinhardt HC, Beleggia F. Functional synapses between small cell lung cancer and glutamatergic neurons. bioRxiv doi.org/10.1101/2023.01.19.524045
- Bebber CM, Thomas ES, Stroh J, Chen Z, Androulidaki A, Schmitt A, Höhne MN, Stüker L, de Pádua Alves C, Khonsari A, Dammert MA, Parmaksiz F, Tumbrink HL, Beleggia F, Sos ML, Riemer J, George J, Brodesser S, Thomas RK, Reinhardt HC, von Karstedt S. Ferroptosis response segregates small cell lung cancer (SCLC) neuroendocrine subtypes. Nat Commun. 2021 Apr 6;12(1):2048. doi: 10.1038/s41467-021-22336-4.
- Riabinska A, Lehrmann D, Jachimowicz RD, Knittel G, Fritz C, Schmitt A, Geyer A, Heneweer C, Wittersheim M, Frenzel LP, Torgovnick A, Wiederstein JL, Wunderlich CM, Ortmann M, Paillard A, Wößmann W, Borkhardt A, Burdach S, Hansmann ML, Rosenwald A, Perner S, Mall G, Klapper W, Merseburg A, Krüger M, Grüll H, Persigehl T, Wunderlich FT, Peifer M, Utermöhlen O, Büttner R, Beleggia F, Reinhardt HC. ATM activity in T cells is critical for immune surveillance of lymphoma in vivo. Leukemia. 2020 Mar;34(3):771-786. doi: 10.1038/s41375-019-0618-2. Epub 2019 Nov 5. PMID: 31690822.
- Jachimowicz RD, Beleggia F, Isensee J, Velpula BB, Goergens J, Bustos MA, Doll MA, Shenoy A, Checa-Rodriguez C, Wiederstein JL, Baranes-Bachar K, Bartenhagen C, Hertwig F, Teper N, Nishi T, Schmitt A, Distelmaier F, Lüdecke HJ, Albrecht B, Krüger M, Schumacher B, Geiger T, Hoon DSB, Huertas P, Fischer M, Hucho T, Peifer M, Ziv Y, Reinhardt HC, Wieczorek D, Shiloh Y. UBQLN4 Represses Homologous Recombination and Is Overexpressed in Aggressive Tumors. Cell. 2019 Jan 24;176(3):505-519.e22. doi: 10.1016/j.cell.2018.11.024. Epub 2019 Jan 3. PMID: 30612738.
- Erber J, Steiner JD, Isensee J, Lobbes LA, Toschka A, Beleggia F, Schmitt A, Kaiser RWJ, Siedek F, Persigehl T, Hucho T, Reinhardt HC. Dual Inhibition of GLUT1 and the ATR/CHK1 Kinase Axis Displays Synergistic Cytotoxicity in KRAS-Mutant Cancer Cells. Cancer Res. 2019 Oct 1;79(19):4855-4868. doi: 10.1158/0008-5472.CAN-18-3959. Epub 2019 Aug 12. PMID: 31405847.
- Torgovnick A, Heger JM, Liaki V, Isensee J, Schmitt A, Knittel G, Riabinska A, Beleggia F, Laurien L, Leeser U, Jüngst C, Siedek F, Vogel W, Klümper N, Nolte H, Wittersheim M, Tharun L, Castiglione R, Krüger M, Schauss A, Perner S, Pasparakis M, Büttner R, Persigehl T, Hucho T, Herter-Sprie GS, Schumacher B, Reinhardt HC. The Cdkn1aSUPER Mouse as a Tool to Study p53-Mediated Tumor Suppression. Cell Rep. 2018 Oct 23;25(4):1027-1039.e6. doi: 10.1016/j.celrep.2018.09.079. PMID: 30355482.
- Doerr F, George J, Schmitt A, Beleggia F, Rehkämper T, Hermann S, Walter V, Weber JP, Thomas RK, Wittersheim M, Büttner R, Persigehl T, Reinhardt HC. Targeting a non-oncogene addiction to the ATR/CHK1 axis for the treatment of small cell lung cancer. Sci Rep. 2017 Nov 14;7(1):15511. doi: 10.1038/s41598-017-15840-5. PMID: 29138515; PMCID: PMC5686113.
- Knittel G, Rehkämper T, Korovkina D, Liedgens P, Fritz C, Torgovnick A, Al- Baldawi Y, Al-Maarri M, Cun Y, Fedorchenko O, Riabinska A, Beleggia F, Nguyen PH, Wunderlich FT, Ortmann M, Montesinos-Rongen M, Tausch E, Stilgenbauer S, P Frenzel L, Herling M, Herling C, Bahlo J, Hallek M, Peifer M, Buettner R, Persigehl T, Reinhardt HC. Two mouse models reveal an actionable PARP1 dependence in aggressive chronic lymphocytic leukemia. Nat Commun. 2017 Jul 28;8(1):153. doi: 10.1038/s41467-017-00210-6. PMID: 28751718; PMCID: PMC5532225.
- Hatzold J, Beleggia F, Herzig H, Altmüller J, Nürnberg P, Bloch W, Wollnik B, Hammerschmidt M. Tumor suppression in basal keratinocytes via dual non-cell- autonomous functions of a Na,K-ATPase beta subunit. Elife. 2016 May 30;5:e14277. doi: 10.7554/eLife.14277. PMID: 27240166; PMCID: PMC4973367
- Rosin N, Elcioglu NH, Beleggia F, Isgüven P, Altmüller J, Thiele H, Steindl K, Joset P, Rauch A, Nürnberg P, Wollnik B, Yigit G. Mutations in XRCC4 cause primary microcephaly, short stature and increased genomic instability. Hum Mol Genet. 2015 Jul 1;24(13):3708-17. doi: 10.1093/hmg/ddv115. Epub 2015 Apr 3. PMID: 25839420.
- Beleggia F, Li Y, Fan J, Elcioğlu NH, Toker E, Wieland T, Maumenee IH, Akarsu NA, Meitinger T, Strom TM, Lang R, Wollnik B. CRIM1 haploinsufficiency causes defects in eye development in human and mouse. Hum Mol Genet. 2015 Apr 15;24(8):2267-73. doi: 10.1093/hmg/ddu744. Epub 2015 Jan 5. PMID: 25561690; PMCID: PMC4380072
- Keupp K, Beleggia F, Kayserili H, Barnes AM, Steiner M, Semler O, Fischer B, Yigit G, Janda CY, Becker J, Breer S, Altunoglu U, Grünhagen J, Krawitz P, Hecht J, Schinke T, Makareeva E, Lausch E, Cankaya T, Caparrós-Martín JA, Lapunzina P, Temtamy S, Aglan M, Zabel B, Eysel P, Koerber F, Leikin S, Garcia KC, Netzer C, Schönau E, Ruiz-Perez VL, Mundlos S, Amling M, Kornak U, Marini J, Wollnik B. Mutations in WNT1 cause different forms of bone fragility. Am J Hum Genet. 2013 Apr 4;92(4):565-74. doi: 10.1016/j.ajhg.2013.02.010. Epub 2013 Mar 14. PMID: 23499309; PMCID: PMC3617378.