Peifer Lab
Computational Cancer Genomics
The development of massively parallel sequencing has fueled genome-centric cancer research. This technology has enabled us to sequence entire cancer genomes and transcriptomes in a relatively short period of time. However, the data produced by this approach is highly complex and difficult to analyze in order to extract the biologically relevant information. Within our group we are developing new computational methods and analyze sequencing data from tumor specimens.
Our research focus:
We develop and apply computational methods to cancer genome sequencing data. This includes methods call somatic genome alterations such as point mutations, copy number changes, and rearrangements. Furthermore, we develop computational tools for integrative genome analyses, to reconstruct tumor evolution, and recently to analyze single cell sequencing data (both on RNA and DNA).
Our goals:
With the help of these methods, we aim to better understand tumorigenesis, clonal evolution of cancer cells, and therapy resistance.
Our Approach:
With an interdisciplinary team, we provide new computational approaches to analyze and interpret high-throughput cancer sequencing data.
Principal Investigator
Most important publications
Rosswog C, Bartenhagen C, Welte A, Kahlert Y, Hemstedt N, Lorenz W, Cartolano M, Ackermann S, Perner S, Vogel W, Altmüller J, Nürnberg P, Hertwig F, Göhring G, Lilienweiss E, Stütz AM, Korbel JO, Thomas RK, Peifer M*, Fischer M*. Chromothripsis followed by circular recombination drives oncogene amplification in human cancer. Nature Genetics (in press). *corresponding authors
Dentro SC, Leshchiner I, Haase K, Tarabichi M, Wintersinger J, Deshwar AG, Yu K, Rubanova Y, Macintyre G, Demeulemeester J, Vázquez-García I, Kleinheinz K, Livitz DG, Malikic S, Donmez N, Sengupta S, Anur P, Jolly C, Cmero M, Rosebrock D, Schumacher SE, Fan Y, Fittall M, Drews RM, Yao X, Watkins TBK, Lee J, Schlesner M, Zhu H, Adams DJ, McGranahan N, Swanton C, Getz G, Boutros PC, Imielinski M, Beroukhim R, Sahinalp SC, Ji Y, Peifer M, Martincorena I, Markowetz F, Mustonen V, Yuan K, Gerstung M, Spellman PT, Wang W, Morris QD, Wedge DC, Van Loo P; PCAWG Evolution and Heterogeneity Working Group and the PCAWG Consortium. Characterizing genetic intra-tumor heterogeneity across 2,658 human cancer genomes. Cell. 2021 Apr 15;184(8):2239-2254.e39.
Cartolano M, Abedpour N, Achter V, Yang TP, Ackermann S, Fischer M, Peifer M. CaMuS: simultaneous fitting and de novo imputation of cancer mutational signature. Scientific Reports, 2020 Nov 9;10(1):19316.
Klein S*, Quaas A, Noh KW, Cartolano M, Abedpour N, Mauch C, Quantius J, Reinhardt HC, Buettner R, Peifer M*, Helbig D*. Integrative Analysis of Pleomorphic Dermal Sarcomas Reveals Fibroblastic Differentiation and Susceptibility to Immunotherapy. Clinical Cancer Research, 2020 Nov 1;26(21):5638-5645. *corresponding authors
Gerstung M, Jolly C, Leshchiner I, Dentro SC, Gonzalez S, Rosebrock D, Mitchell TJ, Rubanova Y, Anur P, Yu K, Tarabichi M, Deshwar A, Wintersinger J, Kleinheinz K, Vázquez-García I, Haase K, Jerman L, Sengupta S, Macintyre G, Malikic S, Donmez N, Livitz DG, Cmero M, Demeulemeester J, Schumacher S, Fan Y, Yao X, Lee J, Schlesner M, Boutros PC, Bowtell DD, Zhu H, Getz G, Imielinski M, Beroukhim R, Sahinalp SC, Ji Y, Peifer M, Markowetz F, Mustonen V, Yuan K, Wang W, Morris QD; PCAWG Evolution & Heterogeneity Working Group, Spellman PT, Wedge DC, Van Loo P; PCAWG Consortium. The Evolutionary History of 2,658 Cancers. Nature, 2020 Feb;578(7793):122-128.
Ackermann S*, Cartolano M*, Hero B, Welte A, Kahlert Y, Roderwieser A, Bartenhagen C, Walter E, Gecht J, Kerschke L, Volland R, Menon R, Heuckmann JM, Gartlgruber M, Hartlieb S, Henrich KO, Okonechnikov K, Altmüller J, Nürnberg P, Lefever S, de Wilde B, Sand F, Ikram F, Rosswog C, Fischer J, Theissen J, Hertwig F, Singhi AD, Simon T, Vogel W, Perner S, Krug B, Schmidt M, Rahmann S, Achter V, Lang U, Vokuhl C, Ortmann M, Büttner R, Eggert A, Speleman F, O’Sullivan RJ, Thomas RK, Berthold F*, Vandesompele J*, Schramm A*, Westermann F*, Schulte JH*, Peifer M*, Fischer M*. A mechanistic classification of clinical phenotypes in neuroblastoma. Science2018 Dec 7;362(6419):1165-1170. *equal contribution
Cun Y, Yang TP, Achter V, Lang U, Peifer M. Copy-number analysis and inference of subclonal populations in cancer genomes using Sclust. Nature Protocols, 2018 Jun;13(6):1488-1501.
Herling CD, Abedpour N, Weiss J, Schmitt A, Jachimowicz RD, Merkel O, Cartolano M, Oberbeck S, Mayer P, Berg V, Thomalla D, Kutsch N, Stiefelhagen M, Cramer P, Wendtner CM, Persigehl T, Saleh A, Altmüller J, Nürnberg P, Pallasch C, Achter V, Lang U, Eichhorst B, Castiglione R, Schäfer SC, Büttner R, Kreuzer KA, Reinhardt HC, Hallek M, Frenzel LP, Peifer M. Clonal dynamics towards the development of venetoclax resistance in chronic lymphocytic leukemia. Nature Communications, 2018 Feb 20;9(1):727.
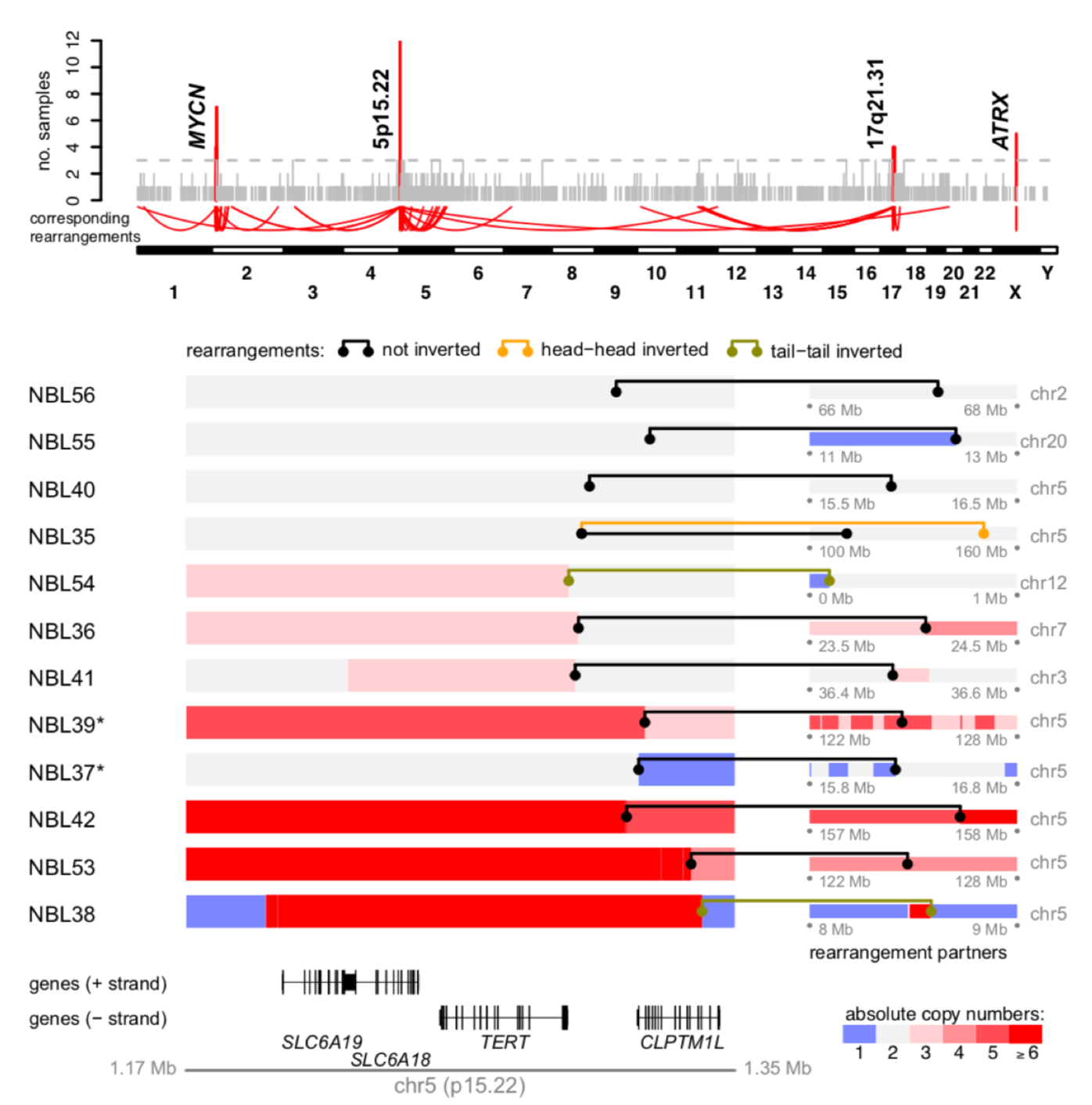
Peifer M*, Hertwig F, Roels F, Dreidax D, Gartlgruber M, Menon R, Krämer A, Roncaioli JL, Sand F, Heuckmann JM, Ikram F, Schmidt R, Ackermann S, Engesser A, Kahlert Y, Vogel W, Altmüller J, Nürnberg P, Thierry-Mieg J, Thierry-Mieg D, Mariappan A, Heynck S, Mariotti E, Henrich KO, Gloeckner C, Bosco G, Leuschner I, Schweiger MR, Savelyeva L, Watkins SC, Shao C, Bell E, Höfer T, Achter V, Lang U, Theissen J, Volland R, Saadati M, Eggert A, de Wilde B, Berthold F, Peng Z, Zhao C, Shi L, Ortmann M, Büttner R, Perner S, Hero B, Schramm A, Schulte JH, Herrmann C, O'Sullivan RJ, Westermann F*, Thomas RK*, Fischer M*. Telomerase activation by genomic rearrangements in high-risk neuroblastoma. Nature, 2015 Oct 29;526(7575):700-4. *corresponding authors
George J, Lim JS, Jang SJ, Cun Y, Ozretic L, Leenders F, Lu X, Fernandez-Cuesta L, Bosco G, Nguyen C, Berg J, Müller C, Dahmen I, Jahchan N, Yang D, Karnezis AN, Vaka D, Torres A, Menon R, Wang MS, Stuetz A, Korbel J, Wilkerson M, Hayes N, Engelmann D, Pützer B, Vlasic I, Seidel D, Pinther B, Schaub B, Becker C, Altmüller J, Yokota J, Khono T, Iwakawa R, Muley T, Peterson I, Chen Y, Soltermann A, Tischler V, Massion P, Zou Y, Wright GM, Russell P, Solomon B, Koch I, Field JK, Muscarella LA, La Torre A, Jakopovic M, Knezevic J, Heiden E, Roz L, Jovanovic D, Kontic M, Thunnissen E, Köhler J, Schuler M, Sanchez-Cespedes M, Salvesen H, Achter V, Bogus M, Schneider P, Zander T, Ansén S, Hallek M, Wolf J, Vingron M, Yatabe Y, Travis WD, Nürnberg P, Reinhardt HC, Perner S, Heukamp LC, Büttner R, Haas SA, Brambilla E, Peifer M*, Sage J*, Thomas RK*. Comprehensive genomic profiling of small cell lung cancer. Nature, 2015 Aug 6;524(7563):47-53. *corresponding authors
Peifer M, Fernández-Cuesta L, Sos ML, George J, Seidel D, Kasper LH, Plenker D, Leenders F, Sun R, Zander T, Menon R, Koker M, Dahmen I, Müller C, Di Cerbo V, Schildhaus HU, Altmüller J, Baessmann I, Becker C, de Wilde B, Vandesompele J, Böhm D, Ansén S, Gabler F, Wilkening I, Heynck S, Heuckmann JM, Lu X, Carter SL, Cibulskis K, Banerji S, Getz G, Park KS, Rauh D, Grütter C, Fischer M, Pasqualucci L, Wright G, Wainer Z, Russell P, Petersen I, Chen Y, Stoelben E, Ludwig C, Schnabel P, Hoffmann H, Muley T, Brockmann M, Engel-Riedel W, Muscarella LA, Fazio VM, Groen H, Timens W, Sietsma H, Thunnissen E, Smit E, Heideman DA, Snijders PJ, Cappuzzo F, Ligorio C, Damiani S, Field J, Solberg S, Brustugun OT, Lund-Iversen M, Sänger J, Clement JH, Soltermann A, Moch H, Weder W, Solomon B, Soria JC, Validire P, Besse B, Brambilla E, Brambilla C, Lantuejoul S, Lorimier P, Schneider PM, Hallek M, Pao W, Meyerson M, Sage J, Shendure J, Schneider R, Büttner R, Wolf J, Nürnberg P, Perner S, Heukamp LC, Brindle PK, Haas S, Thomas RK. Integrative genome analyses identify key somatic driver mutations of small-cell lung cancer. Nature Genetics, Oct;44(10):1104-10.
- Lu X, Thomas RK, Peifer M. CGARS: cancer genome analysis by rank sums. Bioinformatics, 2014 May 1;30(9):1295-6.